wwPDB 2026 News
Contents
05/18/2026
Updated rsync information for PDB Versioned and NextGen archives
Support for the rsync protocol for PDB's Versioned and NextGen archives at RCSB PDB will be discontinued starting June 22.
PDBj will continue to support rsync for these archives at:
- rsync://rsync-versioned.pdbj.org
- rsync://rsync-nextgen.pdbj.org
Https for Versioned and NextGen archive access is available at:
Rsync and http protocols for the recently released Beta archive and the main PDB archive will remain available as before.
04/28/2026
New Syntax for PDB DOIs for Entries Released After July 21, 2027
On July 21, 2027, the wwPDB will fully transition to extended 12-character PDB IDs, the PDBx/mmCIF format, and the re-organized PDB archive. At this point, new 4-character IDs will no longer be issued.
PDB Digital Object Identifiers (DOI) can be used to link to the corresponding wwPDB DOI landing page. DOIs are unique, persistent alphanumeric string assigned to digital objects to provide a permanent, reliable link to their location online. DOIs will not change once assigned.
The DOI syntax for PDB entries after July 20, 2027 will reflect the extended PDB ID.
DOIs for PDB entries deposited after July 20, 2027 (only assigned 12-character PDB IDs)
PDB entries that will only be issued extended PDB IDs (e.g., pdb_1000axyz), will have the same PDB ID in both `_database_2.database_code` and `_database_2.pdbx_database_accession` data items.
These entries will have a new PDB DOI format based on the extended PDB ID, e.g., 10.2210/pdb_1000axyz/pdb.
All 12 characters, including the pdb_ prefix, are included.
loop_
_database_2.database_id
_database_2.database_code
_database_2.pdbx_database_accession
_database_2.pdbx_DOI
PDB pdb_1000axyz pdb_1000axyz 10.2210/pdb_1000axyz/pdb
DOIs for PDB entries deposited before July 21, 2027 (assigned both 4- and 12-character PDB IDs)
The PDB DOI, 4-character PDB ID, and extended 12-character PDB ID are stored in the `database_2` category of the PDBx/mmCIF model file:
loop_
_database_2.database_id
_database_2.database_code
_database_2.pdbx_database_accession
_database_2.pdbx_DOI
PDB 1ABC pdb_00001abc 10.2210/pdb1abc/pdb
For these entries, the PDB DOI format with “pdb” prefix remains unchanged, e.g., 10.2210/pdb1abc/pdb.
For example, the PDB entry with ID 8y9m (and extended PDB ID pdb_00008y9m) has the DOI https://doi.org/10.2210/pdb8y9m/pdb.
Citing PDB DOIs
Many journals use PDB DOIs to link to PDB data from published articles.
PDB DOIs are also used as citations for entries that are not associated with a primary citation.
- PDB DOIs are also used as citations for entries that are not associated with a primary citation.
- Entries that have been issued a 4-character ID (meaning it has been deposited before July 21, 2027) should be cited using the ID and DOI syntax for 4-character IDs (e.g., 10.2210/pdb1abc/pdb). This includes structures with extended IDs created from 4-character IDs using the prefix “pdb_0000” (e.g., “2abc” for pdb_00002abc)
- Entries that have only been issued a 12-character ID (meaning it has been deposited after July 20, 2027) should be cited using the new ID and DOI syntax (e.g., 10.2210/pdb_1000axyz/pdb)
For any further information please contact us at info@wwpdb.org.
Transition Resources
 PDB entries deposited after July 20, 2027 will only be assigned extended PDB IDs and will follow a new DOI syntax.
PDB entries deposited after July 20, 2027 will only be assigned extended PDB IDs and will follow a new DOI syntax.
04/26/2026
July 21, 2027: PDB Beta Archive Will Replace Current Archive
Starting July 21, 2027:
- The PDB Beta Archive will replace the current PDB main archive. The current beta archive URL will point to the main archive.
- PDB entries that have been issued 4-character PDB IDs, including IHM structures in PDB-IHM, will be accessed in this archive using their extended PDB ID that prepends “pdb_0000” to the 4-character ID (e.g., 1ABC becomes pdb_00001abc).
- OneDep will only issue 12-character IDs (prefix pdb_ followed by 8 alphanumeric characters, e.g., pdb_1000axyz).
- New 4-character PDB IDs will not be issued at this point.
- Data deposited after July 21, 2027 will only be released in PDBx/mmCIF and PDBML formats.
- Legacy PDB formatted files will not be generated for these entries.
The PDB Beta Archive is designed to be scalable and extensible. All files in this archive are re-organized using extended PDB IDs (including file naming and directories).
All PDB users, including software developers and journal editors, must transition to this new archive directory structure, PDBx/mmCIF and extended PDB ID format. Start using extended IDs and PDBx/mmCIF data files today.
For any further information please contact us at info@wwpdb.org.
Transition Resources
 PDB Beta Archive Directory Structure
PDB Beta Archive Directory Structure
04/21/2026
Updated Validation Reports for Released PDB and EMDB Entries
All validation reports for released PDB and EMDB entries have been updated and are now available with recalculated percentiles. In addition, validation reports now include new percentile statistics reflecting the state of the PDB archive on December 31, 2025, EDS and MolProbity updates, new Q-score percentile information, and ligand validation improvements.
Ligand validation improvements were made in response to feedback from Global Phasing Ltd., reporting that wwPDB validation software queries of the CSD via Mogul returned insufficient results for such common moieties as adenine, leading to an incorrect rendering of geometric quality in the buster-report-like diagrams. This was traced to a mismatch in aromaticity specifications between wwPDB validation software and Mogul, that has now been updated. In addition, a depiction of non-carbon atoms by grey balls has been added to these diagrams.
The updated reports are accessible from:
A snapshot of the previous version on 20260415 is archived at RCSB PDB and PDBj.
These updated wwPDB validation reports provide an assessment of structure quality using widely accepted standards and criteria, recommended by community experts serving on Validation Task Forces.
Validation reports are provided to depositors through OneDep--the wwPDB portal for validation, deposition and biocuration of structure data. wwPDB partners encourage the use of the stand-alone validation server and the web service API at any time prior to data deposition. Depositors are required to review and accept the reports as part of the data submission process. Validation reports will continue to be developed and improved as we receive recommendations from the expert Validation Task Forces (VTF) for X-ray, NMR, EM, and as we collect feedback from depositors and users.
The wwPDB partners strongly encourage journal editors and referees to request reports from authors as part of the manuscript submission and review process, as already required by Nature, eLife, The Journal of Biological Chemistry, the International Union of Crystallography (IUCr) journals, FEBS journals, Journal of Immunology and Angew Chem Int Ed Engl. Reports are date-stamped, display the wwPDB logo, and represent standardized wwPDB validation.
Further information and sample validation reports are available.
Your feedback, comments, and questions are welcome at validation@mail.wwpdb.org.
03/30/2026
Poster Prize Awarded at ASBMB
The wwPDB Foundation made an award to Sara Scanlan (James Madison University) for The Structural Mechanism of Malate Dehydrogenases and Support for the Reciprocating Mechanism of Catalysis for an outstanding student presentation at the 2026 meeting of the American Society for Biochemistry and Molecular Biology (ASBMB; March 7–10, 2026; National Harbor, MD).
Many thanks to the ASBMB organizers and poster prize judges for making this award possible.
 Christine Zardecki (wwPDB Foundation) and Sara Scanlan (James Madison University)
Christine Zardecki (wwPDB Foundation) and Sara Scanlan (James Madison University)The Structural Mechanism of Malate Dehydrogenases and Support for the Reciprocating Mechanism of Catalysis
Sara Scanlan, Ellis Bell, Christopher Berndsen
James Madison University, Harrisonburg, Virginia
The wwPDB Foundation was established in 2010 to raise funds in support of the outreach activities of the wwPDB. The Foundation raised funds to help support PDB50 events, workshops, and educational publications. The Foundation is chartered as a 501(c)(3) entity exclusively for scientific, literary, charitable, and educational purposes.
The wwPDB Foundation is grateful for our industrial sponsors:
Individual sponsorships are also available.
Consider supporting the next 50 years of PDB's spirit of openness, cooperation, and education with a donation to the wwPDB Foundation.
03/16/2026
Register for PDB Workshops at IUCr2026
wwPDB will be part of the Twenty-Seventh Congress and General Assembly of the International Union of Crystallography (IUCr2026) that will be held August 11-18, 2026 in Calgary, Alberta (Canada).
Register now to join the wwPDB for free, half-day and hands-on workshops that will be held just before the Congress begins. Depositors should register for the workshops during the registration process; discounted pricing for early registration for the overall meeting ends March 31, 2026.
Enhancing PDB Deposition and Validation Practices (afternoon session on Monday, August 10, 2026)
 Pre-meeting workshop: OneDep
Pre-meeting workshop: OneDepThis workshop will provide participants with essential knowledge and skills to easily prepare PDB depositions, evaluate structure quality, and utilize OneDep features to streamline and optimize deposition processes.
Through short lectures and hands-on practical sessions, participants will gain proficiency in making successful PDB depositions using available tools for preparation of necessary data and metadata files consistent with OneDep requirements of the wwPDB OneDep system for complete deposition, rigorous validation, and expert biocuration. Participants will develop a comprehensive understanding of information presented within wwPDB validation reports and the importance in assessing data quality. Practical insights will be shared, enabling participants to confidently undertake and complete successful PDB submissions.
Workshop attendees will also become proficient in leveraging various features and tools integrated into the structure determination software packages and OneDep software system, including automatic initiation of new depositions using application programming interface (API) or using traditional web interface via ORCiD to create deposition sessions, and validating deposited data.
Enabling PDB Deposition of Integrative/Hybrid Methods Structures Determined using 3D Electron Microscopy plus Chemical Crosslinking and Other Complementary Approaches with the PDB-IHM System (afternoon session on Tuesday, August 11, 2026)
 Pre-meeting workshop: IHM Deposition
Pre-meeting workshop: IHM DepositionThis workshop will enable structural biologists to learn about the advantages of using the PDB-IHM system to make complete depositions of 3DEM/Xlinking-MS-based IHM structures and other types of integrative structures to the PDB.
Structures of complex macromolecular assemblies are increasingly determined using integrative or hybrid methods (IHM), wherein complementary experimental and computational techniques are employed. In addition to traditional structure determination methods such as macromolecular crystallography, NMR spectroscopy, and three-dimensional electron microscopy (3DEM), complementary techniques (e.g., chemical crosslinking-mass spectrometry (Xlinking-MS), small-angle scattering, Förster resonance energy transfer, and chemical shift perturbation without NOE restraints) frequently contribute to integrative structure determination.
PDB-IHM, part of the PDB Core Archive, enables deposition, validation, biocuration, preservation, and dissemination of IHM structures. PDB-IHM infrastructure is supported by the IHMCIF data standard (an extension of the PDBx/mmCIF data standard underpinning the PDB) and software tools that ensure FAIR (findable, accessible, interoperable, and reusable) availability of integrative structural biology data. Importantly, integrative structures processed through PDB-IHM receive PDB accession codes and are archived alongside “traditional” single-method experimental structures in the PDB.
The primary goal of this workshop is to facilitate deposition of IHM structures using PDB-IHM functionality, thereby eliminating barriers to fully archiving these structures in the PDB and preventing data loss. Given the predominance of combined 3DEM/Xlinking-MS-based integrative structures, the workshop will highlight these use cases. Through a combination of short lectures and hands-on practical sessions, participants will learn about (a) IHM data deposition requirements, standards, and practices, (b) pre-deposition file preparation, (c) deposition process and workflows, (d) depositing structures through the web interface, (e) submitting multiple structures using an API, and (f) interpreting the information presented in wwPDB validation reports.
Visit IUCr 2026 for complete details about this structural science meeting.
Depositors should register for the workshops during the registration process; early registration ends March 31, 2026.
03/12/2026
Call for Targets: CASP17 (2026)
Advancing the Frontiers of Structural Biology
The Critical Assessment of Structure Prediction (CASP) experiments, held biennially, have recently witnessed a revolution in modeling accuracy driven by deep learning. In 2018, for the first time, the folds of most proteins were correctly computed [1]; in 2020, the accuracy of many computed protein structures rivaled that of experimental ones [2]; and in 2022, there was an enormous increase in the accuracy of computed protein complexes [3].
However, results from CASP16 (2024) suggest a performance plateau [4] in some key areas. As we look toward CASP17 in 2026, our primary goal is to catalyze breakthroughs in areas where deep learning has yet to deliver and where success has major practical implications.
We Need Your Targets
CASP is only possible through the generous participation of the experimental community. A total of 1,300 targets have been obtained over the previous sixteen CASP rounds. CASP is now more tightly focused on areas where deep learning methods are not yet adequate and where there is major applied significance to success. This tighter focus makes obtaining good target sets more challenging than in the past.
We are requesting targets for the 17th round in the following categories:
- Immune Complexes: A major failure area for current deep learning methods, and one with major applied importance. Two promising approaches for solving this problem have been seen in CASP. To find out whether these or others can succeed we need a rich and varied set of non-homologous targets: antibody-antigen, nanobody-antigen complexes, and T-cell receptor complexes in particular.
- Organic Ligand-Protein Complexes: These structures have obvious importance for the development of new small-molecule drugs. The most recent CASP experiment (2024) revealed that deep learning methods deliver results that often fall short of the experimental accuracy. We seek both: sets of 3D protein-ligand complexes for specific receptors and targets with novel ligand chemistries as well as data on affinity rankings.
- Nucleic Acids & Complexes: Despite claims that deep learning methods have solved the problem of computing nucleic acid structures, CASP16 [5] and a subsequent Kaggle challenge [6] showed that these methods are usually no better than classical approaches and that both fail badly in the absence of homologous structural information. But new deep learning methods are appearing now at a fast pace. Thus, for CASP17 we particularly need non-homologous RNA/DNA structures and protein-nucleic acid complexes.
- Conformational Ensembles: Testing methods for computing ensembles of structures, ranging from a few discrete conformations to semi-disordered states, is a major expansion area in CASP. We need targets with multiple high resolution conformations for assessment in the conventional way. We also seek targets where there are multiple lower resolution experimental datasets available, such as cryo-tomography; SAXS; NMR (RDC, chemical shifts and other data); FRET; and cross-linking.
- Difficult Protein Structures and Complexes: In many cases, current deep learning methods deliver high-accuracy structures for single proteins and complexes. But there are critical weaknesses. To help address these, we need targets in the following areas: Membrane proteins; Proteins and complexes with weak evolutionary information such as those with viral or parasite origin, "shallow" sequence alignments and recently evolved interfaces;
- Large proteins and complexes with complex stoichiometry arrangement of subunits (>1,000 amino acids).
Rule of Thumb: If AlphaFold3 can generate a high-quality model, it is likely not a CASP-grade challenge. If it struggles, we want it.
Submission Guidelines & Deadlines
To maintain the rigor of a "blind" test, experimental data must remain confidential (no papers, preprints, PDB releases, or conference presentations) until after the modeling phase.
- Submission window: Now through July 10, 2026.
- Data delivery: Experimental coordinates/data are required by September 1, 2026 (confidentiality can be maintained post-assessment).
- Incentives: Target providers are invited to co-author papers in the special CASP issue of a scientific journal [7-14].
How to Contribute:
- Web Portal (Preferred): Submit via the CASP17 Target Form.
- Direct Email: Contact us at casp@ucdavis.edu to suggest targets or clarify questions.
- PDB Submission: Designate your structure a CASP target with 8 weeks onhold at OneDep deposition interface.
We look forward to your contributions in making CASP17 a landmark event for structural biology.
The CASP Organizers: John Moult, Krzysztof Fidelis, Andriy Kryshtafovych, Torsten Schwede, Maya Topf
- Kryshtafovych A, Schwede T, Topf M, Fidelis K, Moult J. Critical assessment of methods of protein structure prediction (CASP)-Round XIII. Proteins 2019;87(12):1011-1020.
- Kryshtafovych A, Schwede T, Topf M, Fidelis K, Moult J. Critical assessment of methods of protein structure prediction (CASP)-Round XIV. Proteins 2021;89(12):1607-1617.
- Kryshtafovych A, Schwede T, Topf M, Fidelis K, Moult J. Critical assessment of methods of protein structure prediction (CASP)-Round XV. Proteins 2023;91(12):1539-1549.
- Kryshtafovych A, Schwede T, Topf M, Fidelis K, Moult J. Progress and Bottlenecks for Deep Learning in Computational Structure Biology: CASP Round XVI. Proteins 2026; 94(1):5–14.
- Kretsch RC, Hummer AM, He S, Yuan R, Zhang J, Karagianes T, Cong Q, Kryshtafovych A, Das R. Assessment of Nucleic Acid Structure Prediction in CASP16.Proteins. 2026 Jan;94(1):192-217.
- Lee Y, He S, Oda T, Rao GJ, Kim Y, Kim R, Kim H, Heng CK, Kowerko D, Li H, Nguyen H, Sampathkumar A, Gómez RE, Chen M, Yoshizawa A, Kuraishi S, Ogawa K, Zou S, Paullier A, Zhao B, Chen HL, Hsu TA, Hirano T, Chiu W, Gezelle JG, Haack D, Hong Y, Jadhav S, Koirala D, Kretsch RC, Lewicka A, Li S, Marcia M, Piccirilli J, Rudolfs B, Srivastava Y, Steckelberg AL, Su Z, Toor N, Wang L, Yang Z, Zhang K, Zou J, Baker D, Chen SJ, Demkin M, Favor A, Hummer AM, Joshi CK, Kryshtafovych A, Küçükbenli E, Miao Z, Moult J, Munley C, Reade W, Viel T, Westhof E, Zhang S, Das R. Template-based RNA structure prediction advanced through a blind code competition.
- bioRxiv [Preprint]. 2025 Dec 30:2025.12.30.696949. doi: 10.64898/2025.12.30.696949.
- Kryshtafovych A, Albrecht R, Basle A, Bule P, Caputo AT, Carvalho AL, Chao KL, Diskin R, Fidelis K, Fontes C, Fredslund F, Gilbert HJ, Goulding CW, Hartmann MD, Hayes CS, Herzberg O, Hill JC, Joachimiak A, Kohring GW, Koning RI, Lo Leggio L, Mangiagalli M, Michalska K, Moult J, Najmudin S, Nardini M, Nardone V, Ndeh D, Nguyen TH, Pintacuda G, Postel S, van Raaij MJ, Roversi P, Shimon A, Singh AK, Sundberg EJ, Tars K, Zitzmann N, Schwede T. Target highlights from the first post-PSI CASP experiment (CASP12, May-August 2016). Proteins 2018;86 Suppl 1(Suppl 1):27-50.
- Lepore R, Kryshtafovych A, Alahuhta M, Veraszto HA, Bomble YJ, Bufton JC, Bullock AN, Caba C, Cao H, Davies OR, Desfosses A, Dunne M, Fidelis K, Goulding CW, Gurusaran M, Gutsche I, Harding CJ, Hartmann MD, Hayes CS, Joachimiak A, Leiman PG, Loppnau P, Lovering AL, Lunin VV, Michalska K, Mir-Sanchis I, Mitra AK, Moult J, Phillips GN, Jr., Pinkas DM, Rice PA, Tong Y, Topf M, Walton JD, Schwede T. Target highlights in CASP13: Experimental target structures through the eyes of their authors. Proteins 2019;87(12):1037-1057.
- Alexander LT, Lepore R, Kryshtafovych A, Adamopoulos A, Alahuhta M, Arvin AM, Bomble YJ, Bottcher B, Breyton C, Chiarini V, Chinnam NB, Chiu W, Fidelis K, Grinter R, Gupta GD, Hartmann MD, Hayes CS, Heidebrecht T, Ilari A, Joachimiak A, Kim Y, Linares R, Lovering AL, Lunin VV, Lupas AN, Makbul C, Michalska K, Moult J, Mukherjee PK, Nutt WS, Oliver SL, Perrakis A, Stols L, Tainer JA, Topf M, Tsutakawa SE, Valdivia-Delgado M, Schwede T. Target highlights in CASP14: Analysis of models by structure providers. Proteins 2021;89(12):1647-1672.
- Kretsch RC, Andersen ES, Bujnicki JM, Chiu W, Das R, Luo B, Masquida B, McRae EKS, Schroeder GM, Su Z, Wedekind JE, Xu L, Zhang K, Zheludev IN, Moult J, Kryshtafovych A. RNA target highlights in CASP15: Evaluation of predicted models by structure providers. Proteins 2023;91(12):1600-1615.
- Alexander LT, Durairaj J, Kryshtafovych A, Abriata LA, Bayo Y, Bhabha G, Breyton C, Caulton SG, Chen J, Degroux S, Ekiert DC, Erlandsen BS, Freddolino PL, Gilzer D, Greening C, Grimes JM, Grinter R, Gurusaran M, Hartmann MD, Hitchman CJ, Keown JR, Kropp A, Kursula P, Lovering AL, Lemaitre B, Lia A, Liu S, Logotheti M, Lu S, Markusson S, Miller MD, Minasov G, Niemann HH, Opazo F, Phillips GN, Jr., Davies OR, Rommelaere S, Rosas-Lemus M, Roversi P, Satchell K, Smith N, Wilson MA, Wu KL, Xia X, Xiao H, Zhang W, Zhou ZH, Fidelis K, Topf M, Moult J, Schwede T. Protein target highlights in CASP15: Analysis of models by structure providers. Proteins 2023;91(12):1571-1599.
- Alexander LT, Follonier OM, Kryshtafovych A, Abesamis K, Bibi-Triki S, Box HG, Breyton C, Bringel F, Carrique L, d'Acapito A, Dong G, DuBois R, Fass D, Fiesco JM, Fox DR, Grimes JM, Grinter R, Jenkins M, Kamyshinsky R, Keown JR, Lackner G, Lammers M, Liu S, Lovering AL, Malinauskas T, Masquida B, Palm GJ, Siebold C, Su T, Zhang P, Zhou ZH, Fidelis K, Topf M, Moult J, Schwede T. Protein Target Highlights in CASP16: Insights From the Structure Providers. Proteins. 2026 Jan;94(1):25-50. doi: 10.1002/prot.70025.
- Tosstorff A, Rudolph MG, Benz J, Kuhn B, Kramer C, Sharpe M, Huang CY, Metz A, Hazemann J, Ritz D, Sweeney AM, Gilson MK. The CASP 16 Experimental Protein-Ligand Datasets. Proteins. 2026 Jan;94(1):79-85. doi: 10.1002/prot.70053.
- Kretsch RC, Albrecht R, Andersen ES, Chen HA, Chiu W, Das R, Gezelle JG, Hartmann MD, Höbartner C, Hu Y, Jadhav S, Johnson PE, Jones CP, Koirala D, Kristoffersen EL, Largy E, Lewicka A, Mackereth CD, Marcia M, Nigro M, Ojha M, Piccirilli JA, Rice PA, Shin H, Steckelberg AL, Su Z, Srivastava Y, Wang L, Wu Y, Xie J, Zwergius NH, Moult J, Kryshtafovych A. Functional Relevance of CASP16 Nucleic Acid Predictions as Evaluated by Structure Providers. Proteins. 2026 Jan;94(1):51-78. doi: 10.1002/prot.70043.
03/10/2026
Poster Prize Awarded at The Biophysical Society Meeting
The wwPDB Foundation made an award for an outstanding student presentation at the 2026 meeting of The Biophysical Society (February 21-25, 2026; San Francisco, CA).
Many thanks to The Biophysical Society organizers and poster prize judges for making this award possible.
 Haley R. Johnson
Haley R. JohnsonBase Hunting, A New Approach to Base Identification to Determine DNA Sequence from Cryo-EM Density Maps
Haley R. Johnson(1), Matthew L. Baker(2), Jonathan M. Fogg(1), Lynn Zechiedrich(1)
(1) Baylor College of Medicine, Houston, TX, USA
(2) University of Texas Health Science Center, Houston, TX, USA
The wwPDB Foundation was established in 2010 to raise funds in support of the outreach activities of the wwPDB. The Foundation raised funds to help support PDB50 events, workshops, and educational publications. The Foundation is chartered as a 501(c)(3) entity exclusively for scientific, literary, charitable, and educational purposes.
The wwPDB Foundation is grateful for our industrial sponsors. Individual sponsorships are also available.
Consider supporting the next 50 years of PDB's spirit of openness, cooperation, and education with a donation to the wwPDB Foundation.
03/05/2026
PDB Reaches a New Milestone
 The PDB contains more than 250,000 entries
The PDB contains more than 250,000 entries Depositors: Download this image, write the number of structures deposited, and tag us in your photos
Depositors: Download this image, write the number of structures deposited, and tag us in your photosWith this week's update, the PDB archive contains a record 250,441 entries.
Established in 1971, this central, public archive has reached this critical milestone thanks to the efforts of structural biologists throughout the world who contribute their experimentally-determined protein and nucleic acid structure data.
wwPDB data centers support online access to three-dimensional structures of biological macromolecules that help researchers understand many facets of biomedicine, agriculture, and ecology, from protein synthesis to health and disease to biological energy.
The wwPDB passed the 200,000 milestone in 2023. More than 20,000 new structure depositions were submitted to the archive in 2025; PDB data were accessed more than 4.7 billion times in the same year.
Function follows form
In the 1950s, scientists had their first direct look at the structures of proteins and DNA at the atomic level. Determination of these early three-dimensional structures by X-ray crystallography ushered in a new era in biology-one driven by the intimate link between form and biological function. As the value of archiving and sharing these data were quickly recognized by the scientific community, the Protein Data Bank (PDB) was established as the first open access digital resource in all of biology by an international collaboration in 1971 with data centers located in the US and the UK.
Among the first structures deposited in the PDB were those of myoglobin and hemoglobin, two oxygen-binding molecules whose structures were elucidated by Chemistry Nobel Laureates John Kendrew and Max Perutz. With this week's regular update, the PDB welcomes 266 new structures into the archive. These structures join others vital to drug discovery, bioinformatics and education.
The PDB is growing rapidly, increasing in size by ~160% since 2011 (doubling in size every 6-8 years). In 2022, an average of 275 new structures were released to the scientific community each week. The resource is accessed hundreds of millions of times annually by researchers, students, and educators intent on exploring how different proteins are related to one another, to clarify fundamental biological mechanisms and discover new medicines.
More than Twenty Years of Collaboration
Since its inception, the PDB has been a community-driven enterprise, evolving into a mission critical international resource for biological research. The wwPDB partnership was established in July 2003 with PDBe, PDBj, and RCSB PDB. Today, the collaboration includes partners BMRB (joined in 2006) and EMDB (2021).
The wwPDB ensures that these valuable atomic coordinate and experimental supporting data are securely stored, expertly managed, and made freely available for the benefit of scientists and educators around the globe. wwPDB data centers work closely with community experts to define deposition and annotation policies, resolve data representation issues, and implement community validation standards. In addition, the wwPDB works to raise the profile of structural biology with increasingly broad audiences.
Each structure submitted to the archive is carefully curated by wwPDB staff before release. New depositions are checked and enhanced with value-added annotations and linked with other important biological data to ensure that atomic coordinate and experimental supporting data are discoverable and interpretable by users with a wide range of backgrounds and interests.
wwPDB eagerly awaits the next 50,000 structures and the invaluable knowledge these new data will bring.
02/10/2026
New PDB Beta Archive Available for Testing
By 2028 4-character PDB IDs (e.g. 1abc) will be fully allocated. After that, all new entries will be assigned only extended PDB IDs.
The new extended PDB ID format will be 12 characters, which includes a prefix pdb_ followed by 8 alphanumeric characters, e.g. pdb_1000axyz. This new ID format will enable text mining detection of PDB entries in the published literature and allow for more informative and transparent delivery of revised data files. When submitting extended PDB IDs to journals and citing extended PDB IDs in manuscripts, all 12 characters including prefix pdb_ should be provided.
A PDB Beta Archive is now available to help community adopt extended PDB ID and PDBx/mmCIF format during the transition phase. All files at this archive are re-organized with extended PDB ID (including file naming and directories) at entry level, mirroring the same data organization of the PDB Versioned Archive.
All data files for a particular entry are stored in a single directory, labeled based on a two-character hash generated from the penultimate two characters of the PDB code, i.e., https://files-beta.wwpdb.org/pub/wwpdb/pdb/data/entries///. The two-letter hash will be based on the second and third characters from the last character. For example, PDB entry pdb_1abc5678 will be under /67/. This will maintain consistency with the current PDB archive: PDB entry 1abc is under /ab.
File naming is standardized such that the file type is used for the extension.
For example, file naming is changed from r116dsf.ent.gz to pdb_0000116d-sf.cif.gz for the structure factor file and from pdb318d.ent.gz to pdb_0000318d.pdb.gz for the legacy PDB formatted coordinate file.
When four character PDB IDs are about to be consumed, this PDB Beta Archive will replace the current PDB Archive (expected to be around mid-2027) and entries with extended PDB IDs issued are not compatible with PDB format. wwPDB encourages scientific journals, PDB community and users to transition to PDBx/mmCIF format and adopt new PDB ID format as earlier as possible.
For any further information please contact us at info@wwpdb.org.
 New PDB beta archive available for testing in support of extended PDB IDs
New PDB beta archive available for testing in support of extended PDB IDs
01/19/2026
EMDB Surpasses 10,000 Entries in a Year
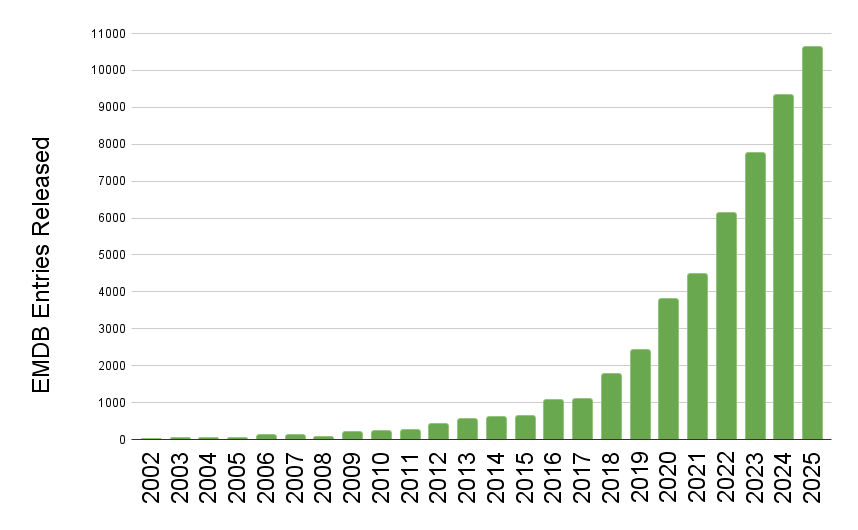 EMDB released entry growth by year
EMDB released entry growth by yearThe wwPDB has reached a historic milestone, releasing 10,000 EMDB entries in a single year.
The rate of release of 3DEM data to the public continues at an extraordinary pace, for the first time the wwPDB has seen the release of 10,000 entries from the EMDB in just a single year. To put this in context, in 2019 the EMDB stood at 10,000 entries in total, after 17 years of growth. Now in 2025 due to the current volume of data release, the archive has surpassed 50,000 total entries.
This rapid growth is a testament to the hard work of the community which the wwPDB endeavors to support. It also highlights the ongoing challenge the community is facing with ever more data created requiring streamlined deposition, processing, curation, peer-review, and sustainable & scalable archiving practices. We are excited to continue to be part of structural biology history.
01/12/2026
wwPDB Shared Responsibilities on Data Processing
Depositions are automatically distributed to wwPDB deposition centers geographically.
Data processing responsibilities are shared among wwPDB partner sites:
- RCSB PDB is responsible for PDB and EMDB data coming from the Americas
- PDBe and EMDB are responsible for PDB and EMDB data from Europe (except Belarus and Russian Federation) and Africa
- PDBj is responsible for PDB and EMDB data from Asia regions (except China), Belarus, Russian Federation and Oceania
- PDBc is responsible for PDB and EMDB data from China
- BMRB is responsible for BMRB data from the Americas, Africa, and Europe
- BMRBj is responsible for BMRB data from Asia
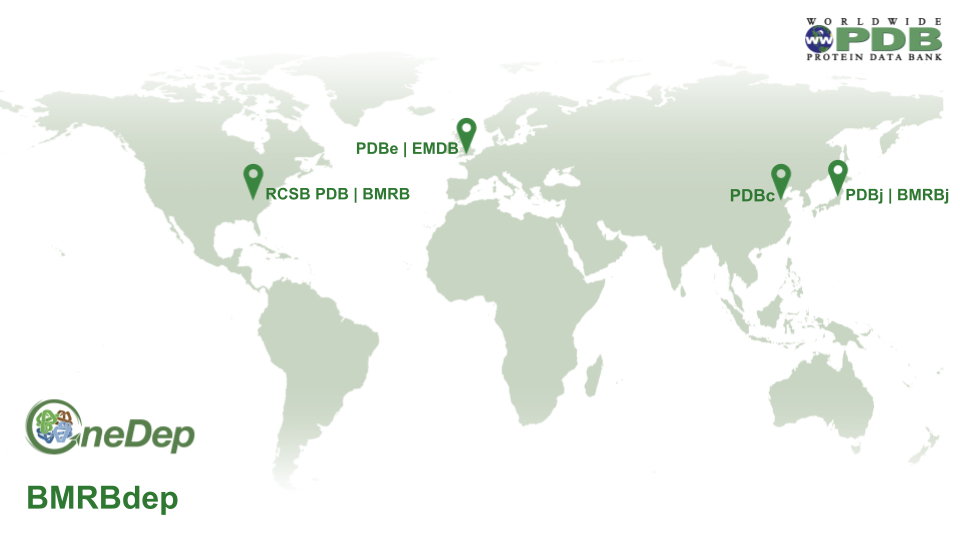 Data processing distribution map
Data processing distribution map
01/11/2026
Register for PDB Workshops at IUCr2026
wwPDB will be part of the Twenty-Seventh Congress and General Assembly of the International Union of Crystallography (IUCr2026) that will be held August 11-18, 2026 in Calgary, Alberta (Canada).
Register now to join the wwPDB for free, half-day and hands-on workshops that will be held just before the Congress begins.
Enhancing PDB Deposition and Validation Practices (afternoon session on Monday, August 10, 2026)
 Pre-meeting workshop: OneDep
Pre-meeting workshop: OneDepThis workshop will provide participants with essential knowledge and skills to easily prepare PDB depositions, evaluate structure quality, and utilize OneDep features to streamline and optimize deposition processes.
Through short lectures and hands-on practical sessions, participants will gain proficiency in making successful PDB depositions using available tools for preparation of necessary data and metadata files consistent with OneDep requirements of the wwPDB OneDep system for complete deposition, rigorous validation, and expert biocuration. Participants will develop a comprehensive understanding of information presented within wwPDB validation reports and the importance in assessing data quality. Practical insights will be shared, enabling participants to confidently undertake and complete successful PDB submissions.
Workshop attendees will also become proficient in leveraging various features and tools integrated into the structure determination software packages and OneDep software system, including automatic initiation of new depositions using application programming interface (API) or using traditional web interface via ORCiD to create deposition sessions, and validating deposited data.
Enabling PDB Deposition of Integrative/Hybrid Methods Structures Determined using 3D Electron Microscopy plus Chemical Crosslinking and Other Complementary Approaches with the PDB-IHM System (afternoon session on Tuesday, August 11, 2026)
 Pre-meeting workshop: IHM Deposition
Pre-meeting workshop: IHM DepositionThis workshop will enable structural biologists to learn about the advantages of using the PDB-IHM system to make complete depositions of 3DEM/Xlinking-MS-based IHM structures and other types of integrative structures to the PDB.
Structures of complex macromolecular assemblies are increasingly determined using integrative or hybrid methods (IHM), wherein complementary experimental and computational techniques are employed. In addition to traditional structure determination methods such as macromolecular crystallography, NMR spectroscopy, and three-dimensional electron microscopy (3DEM), complementary techniques (e.g., chemical crosslinking-mass spectrometry (Xlinking-MS), small-angle scattering, Förster resonance energy transfer, and chemical shift perturbation without NOE restraints) frequently contribute to integrative structure determination.
PDB-IHM, part of the PDB Core Archive, enables deposition, validation, biocuration, preservation, and dissemination of IHM structures. PDB-IHM infrastructure is supported by the IHMCIF data standard (an extension of the PDBx/mmCIF data standard underpinning the PDB) and software tools that ensure FAIR (findable, accessible, interoperable, and reusable) availability of integrative structural biology data. Importantly, integrative structures processed through PDB-IHM receive PDB accession codes and are archived alongside “traditional” single-method experimental structures in the PDB.
The primary goal of this workshop is to facilitate deposition of IHM structures using PDB-IHM functionality, thereby eliminating barriers to fully archiving these structures in the PDB and preventing data loss. Given the predominance of combined 3DEM/Xlinking-MS-based integrative structures, the workshop will highlight these use cases. Through a combination of short lectures and hands-on practical sessions, participants will learn about (a) IHM data deposition requirements, standards, and practices, (b) pre-deposition file preparation, (c) deposition process and workflows, (d) depositing structures through the web interface, (e) submitting multiple structures using an API, and (f) interpreting the information presented in wwPDB validation reports.
Visit IUCr 2026 for complete details about this structural science meeting.
Depositors should register for the workshops during the registration process; early registration ends March 31, 2026.
01/07/2026
Time-stamped Copies of PDB and EMDB Archives
A snapshot of the PDB Core archive (https://files.wwpdb.org, https://s3.rcsb.org) as of January 01, 2026 has been added to https://s3snapshots.rcsb.org (AWS), snapshots.rcsb.org (rsync -rlpy -a -v --delete snapshots.rcsb.org:: .), and ftp://snapshots.pdbj.org. Snapshots have been archived annually since 2005 to provide readily identifiable data sets for research on the PDB archive.
The directory 20260101 includes the 246,905 experimentally-determined structure and experimental data available at that time. Atomic coordinate and related metadata are available in PDBx/mmCIF, PDB, and XML file formats. The date and time stamp of each file indicates the last time the file was modified. The snapshot of PDB Core Archive is 1,583 GB.
A snapshot of the EMDB Core archive (ftp://ftp.ebi.ac.uk/pub/databases/emdb/) as of January 01, 2026 can be found in https://ftp.ebi.ac.uk/pub/databases/emdb_vault/20260101/ and ftp://snapshots.pdbj.org/20260101/. The snapshot includes 52,943 3DEM entries. The Snapshot of the EMDB archive is 28.2 TB in size.